Lung adenocarcinoma (LUAD) is the most common histological subtype of lung cancer, with an average 5-year survival rate of 15%. In contrast, the pre-invasive stages of LUAD, such as adenocarcinoma in situ (AIS) and minimally invasive adenocarcinoma (MIA), are associated with a nearly 100% recurrence-free survival rate after surgical resection.
AIS is defined as a≤3 cm adenocarcinoma lacking invasion, while MIA is a≤3 cm adenocarcinoma with≤5mm invasion. Although some focused studies have identified mutations in lung cancer drivers in AIS and MIA, there remains a lack of deep insight into the molecular events driving progression of these lesions to invasive LUAD. People have little knowledge of the genomic and immune profiles of pre-invasive stages of LUAD, such as AIS and MIA pathogenesis.
Recently, a research titled “Genomic and immune profiling of pre-invasive lung adenocarcinoma” was published in Nature Communications. The research was jointly conducted by two teams led by Prof. Chen Haiquan from Fudan University Shanghai Cancer Center (FUSCC) and Matthew Meyerson from Harvard Medical School.
The researchers performed whole-exome sequencing (WES) and RNA-sequencing (RNA-seq) on tumor and matched adjacent normal tissue of 24 AIS, 74 MIA, and 99 invasive LUAD samples. They identified EGFR, RBM10, BRAF, ERBB2, TP53, KRAS, MAP2K1 and MET as significantly mutated genes in the pre/minimally invasive group. Classes of genome alterations that increase in frequency during the progression to malignancy were revealed, including mutations in TP53, arm-level copy number alterations and HLA loss of heterozygosity. The research found that immune infiltration was correlated with copy number alterations of chromosome arm 6p, suggesting a link between arm-level events and the tumor immune environment.
The research identified eight significantly mutated genes in AIS and MIA specimens, including EGFR, RBM10, BRAF, ERBB2, TP53, KRAS, MAP2K1, and MET, all previously reported as recurrently mutated in LUAD from The Cancer Genome Atlas (TCGA) cohort. EGFR, TP53, RB1 and KRAS were significantly mutated in the tested LUAD cases (Fig. 1a, b).Amplified regions that included MDM2, MYC, TERT, KRAS, NKX2-1, and CDK6 were observed in the AIS or MIA samples (Fig. 1c). Novel amplifications of RIT1 were identified in the FUSCC LUAD cohort. RNA-seq analysis revealed a RET fusion in an MIA sample (Fig. a), and ALK and ROS1 fusions in LUAD (Fig. 1b). When testing significantly mutated genes, TP53 mutations were the most enriched alteration in the invasive stage (38%) compared to pre/minimally invasive stages (6%), followed by EGFR and RB1 mutations (Fig. 1d). Tumor mutation burden (TMB) in stage I LUAD was significantly higher than that in AIS and MIA, the APOBEC signature was also higher, and arm-level copy-number alteration (CNA) was more common.
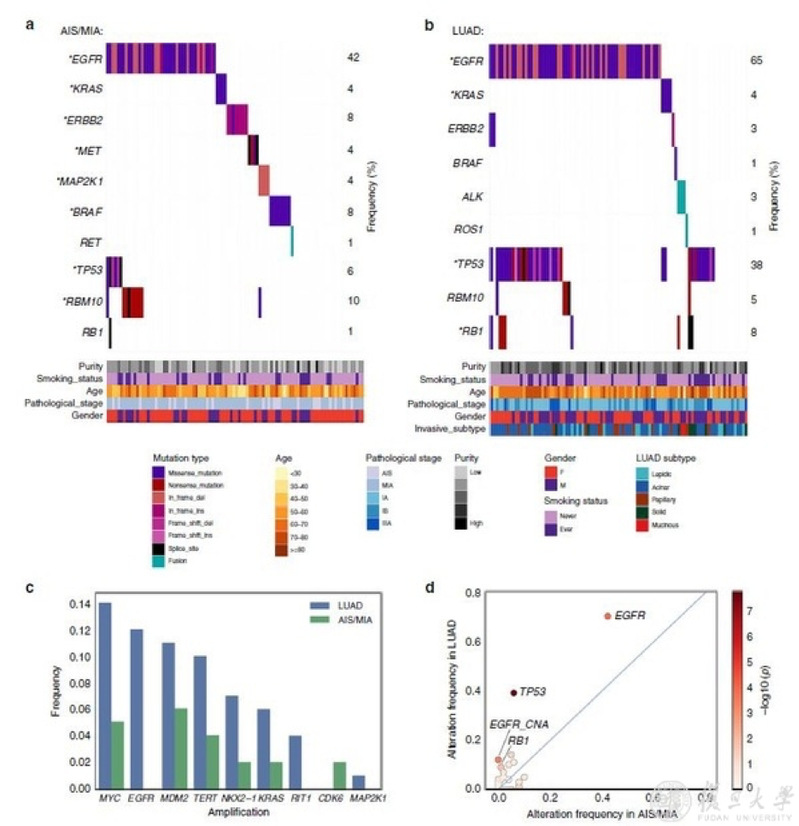
Fig. 1 Somatic alterations in pre-invasive and invasive lung adenocarcinomas.
Loss of human leukocyte antigen (HLA) alleles has been identified as a potential immune escape mechanism in lung cancers. The researchers noted HLA loss of heterozygosity (LOH) in 3.1% of AIS/MIA and 16.7% of LUAD specimens. The significantly increased frequency of HLA LOH in the invasive group compared to the pre-invasive group (Fisher’s exact test, p < 0.01) suggested the potential role of loss of HLA alleles during tumor development.
Approximately 60% of the HLA LOH events in LUAD were related to loss of chromosome 6p. Interestingly, 6p gain was discovered to be significantly anticorrelated with T cell abundance (Mann–Whitney U test, p = 0.038, Fig. 2), and this trend was also observed when analyzing B cell infiltration in correlation with 6p CNA. A subsequent test was done to study the correlation of immune infiltration with large-scale chromosome alterations, using samples from the TCGA LUAD cohort.
The researchers observed the most significant correlation of leukocyte fraction with chromosome 6p CNA (p = 0.0030, coef. = −0.74, 95% CI: −1.23 to −0.25), followed by 1q (p = 0.0033, coef. = −0.60, 95% CI: −1 to −0.2) and 19p CNA (p = 0.0047, coef. = 0.53, 95% CI: 0.16 to 0.9), after controlling for TMB and the degree of overall aneuploidy (Fig. 2c, d). 6p and 1q CNA showed significantly increased frequency from AIS/MIA to LUAD in the FUSCC cohort (Fisher’s exact test, p < 0.001).
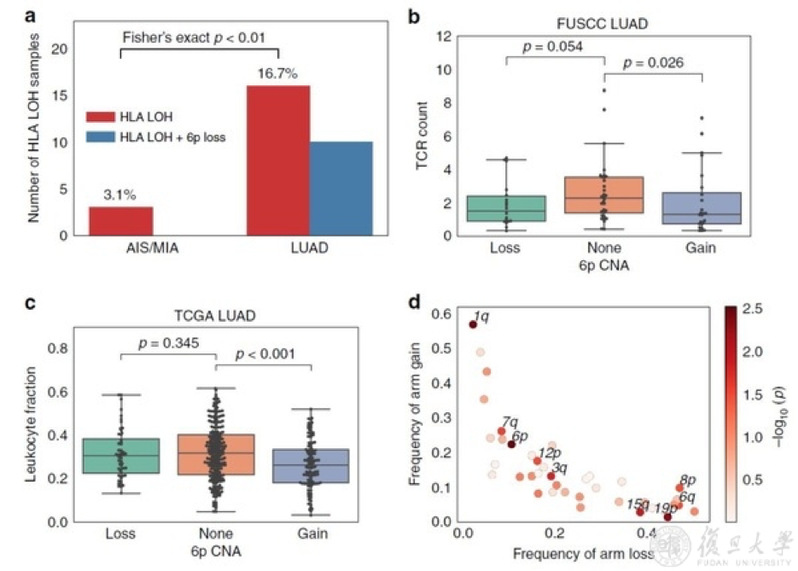
Fig 2. Tumor immune environment in association with arm-level CNA
The research interrogated the genomic and immune features of pre/minimally invasive lung cancers. 71% of AIS and MIA patients carried at least one mutation in previously identified cancer genes in the RTK/RAS/RAF pathway, similar to the oncogenic driver events found in LUAD.
It also revealed TP53 as a key mediator in the invasiveness of lung cancer. Previous studies in Barrett’s esophagus suggested that TP53 occurred early in esophageal adenocarcinoma precursors followed by oncogenic amplifications. TP53 was also frequently mutated in lung carcinoma in situ, which is the precursor form of squamous cell carcinoma.
The research has shown the high frequency of oncogenic driver mutations, but low frequency of TP53 mutations in the LUAD precursors. Previous studies have suggested the functional association of TP53 mutations with invasive potential in cancers. This study also demonstrated a strong association of TP53 mutations with aneuploidy, in line with recent work from TGCA.
Given previous reports of aneuploidy in association with decreased immune infiltration, the research data raised the possibility that copy-number changes in specific chromosomes might influence the tumor microenvironment, thus providing new insights into the biology of lung pre-malignancy, with implications for disease monitoring and prognosis, and future therapeutic intervention.
Full text link:https://www.nature.com/articles/s41467-019-13460-3.pdf





